Dynamics Studies of Intrinsically Disordered Protein by NMR
Potential applications from field-cycling protein NMR
In Biological reactions, many intrinsically disorder proteins (IDPs) play crucial roles. Even though they do not behave as conformational proteins to have certain structures to select/interact with their substracts, their interactions in many pathways are dominated by altering dynamics, some even conformation changes, from disordered to order.
Since IDPs have no solid structures, their studies are restricted in crystal-related techniques. Although the NMR technique has no such restriction, the challenge of NMR is to have well-dispersion spectra to make sufficient resolution on chemical shift assignments. Nowadays, the development of NMR has successfully provided sufficient spectral resolution to offer unique opportunities for conformation and dynamics studies of IDPs. During the last few decades, NMR chemical shift, residual dipolar coupling (RDC), and hydrogen exchange rates are developed to evaluate local transient secondary structural elements with atomic resolution. Furthermore, paramagnetic relaxation enhancement (PRE) investigates long-range contacts.
In addition to those techniques in NMR, field-Cycling NMR spin relaxation study could intensively observe IDP dynamics due to its rapid correlation time. Low field relaxation brings more information on IDP than high-field spin relaxation. The relaxation dispersion curves of IDP could reveal dynamics correlation on dipolar mechanism because the measurable relaxation rate is comparably lower than structural proteins. However, the spectral resolution of the fixed-field relaxation study is restricted by the operating magnetic field. The High-field field-cycling NMR technique has broken this barrier of limited resolution in low-field spin relaxation study by applying a high-magnetic spectrometer as a "spectrum-resolution provider". Investigation IDP dynamics could definitely be benefited by such a high-field field-cycling NMR technique.
Related Literature
NMR contributions to structural dynamics studies of intrinsically disordered proteins
Robert Konrat, JMR 2014, 241, 74-85.
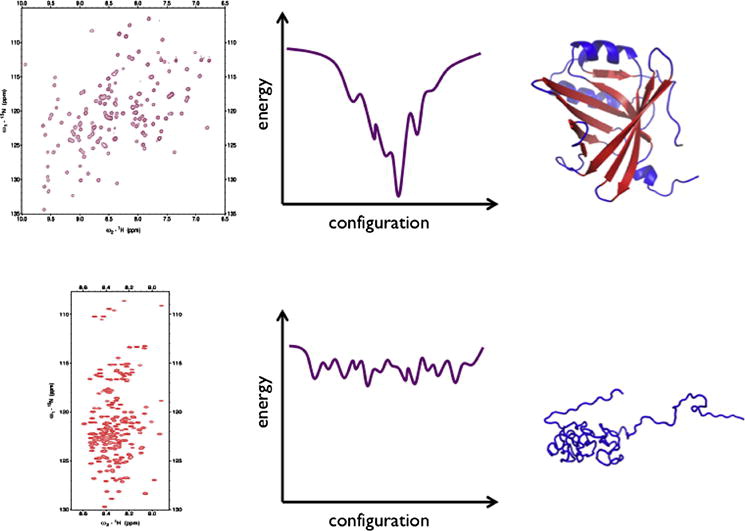
Intrinsically disordered proteins (IDPs) are characterized by substantial conformational plasticity. Given their inherent structural flexibility X-ray crystallography is not applicable to study these proteins. In contrast, NMR spectroscopy offers unique opportunities for structural and dynamic studies of IDPs. The past two decades have witnessed significant development of NMR spectroscopy that couples advances in spin physics and chemistry with a broad range of applications. This article will summarize key advances in basic physical-chemistry and NMR methodology, outline their limitations and envision future research and development directions.
Characterization of intrinsically disordered proteins and their dynamic complexes: From in vitro to cell-like environments
Sigrid Milles, Nicola Salvi, Martin Blackledge and Malene Ringkjøbing Jensen
Progress in Nuclear Magnetic Resonance Spectroscopy 2018 109, 79-100
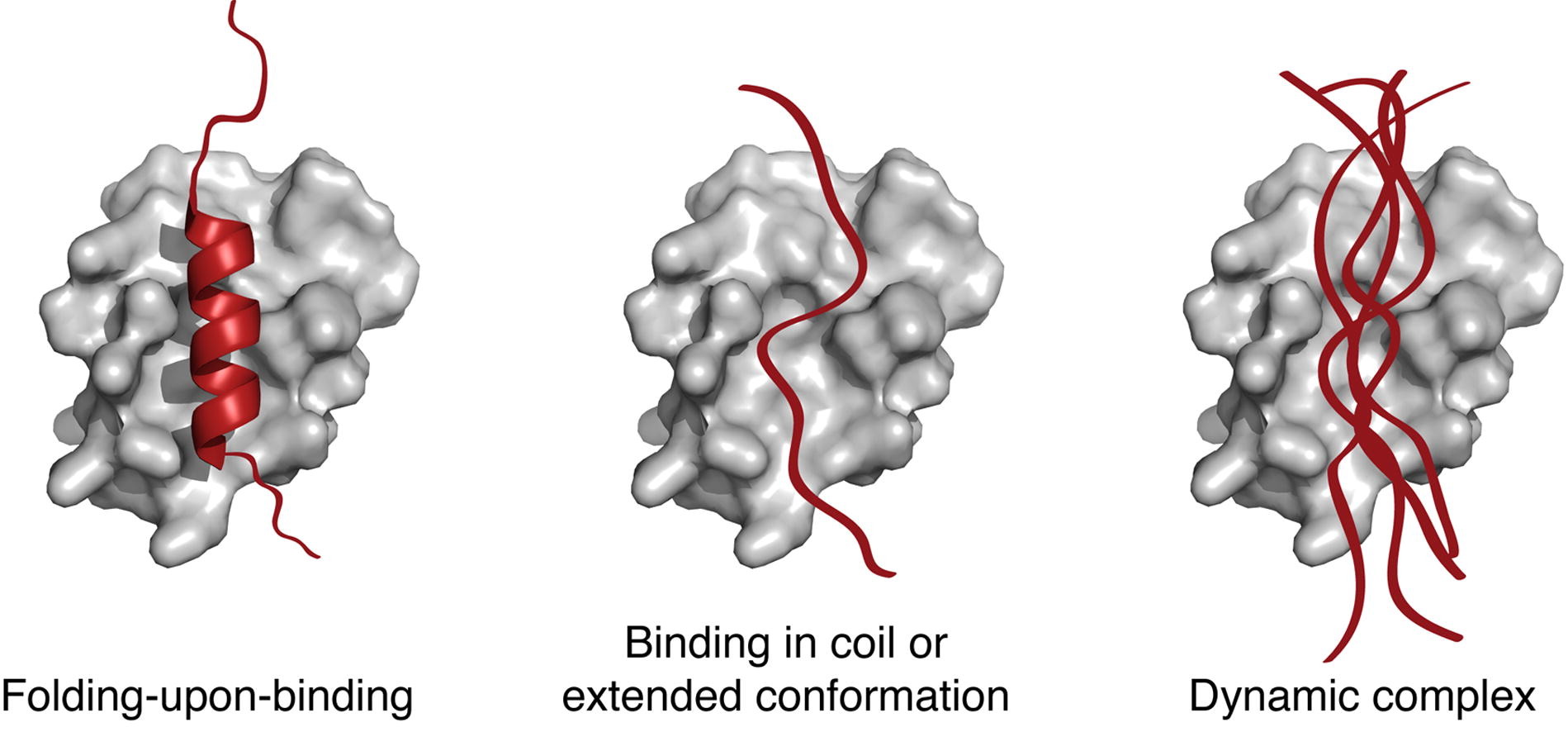
Over the last two decades, it has become increasingly clear that a large fraction of the human proteome is intrinsically disordered or contains disordered segments of significant length. These intrinsically disordered proteins (IDPs) play important regulatory roles throughout biology, underlining the importance of understanding their conformational behavior and interaction mechanisms at the molecular level. Here we review recent progress in the NMR characterization of the structure and dynamics of IDPs in various functional states and environments. We describe the complementarity of different NMR parameters for quantifying the conformational propensities of IDPs in their isolated and phosphorylated states, and we discuss the challenges associated with obtaining structural models of dynamic protein-protein complexes involving IDPs. In addition, we review recent progress in understanding the conformational behavior of IDPs in cell-like environments such as in the presence of crowding agents, in membrane-less organelles and in the complex environment of the human cell.